Molecular Markers Database Overview
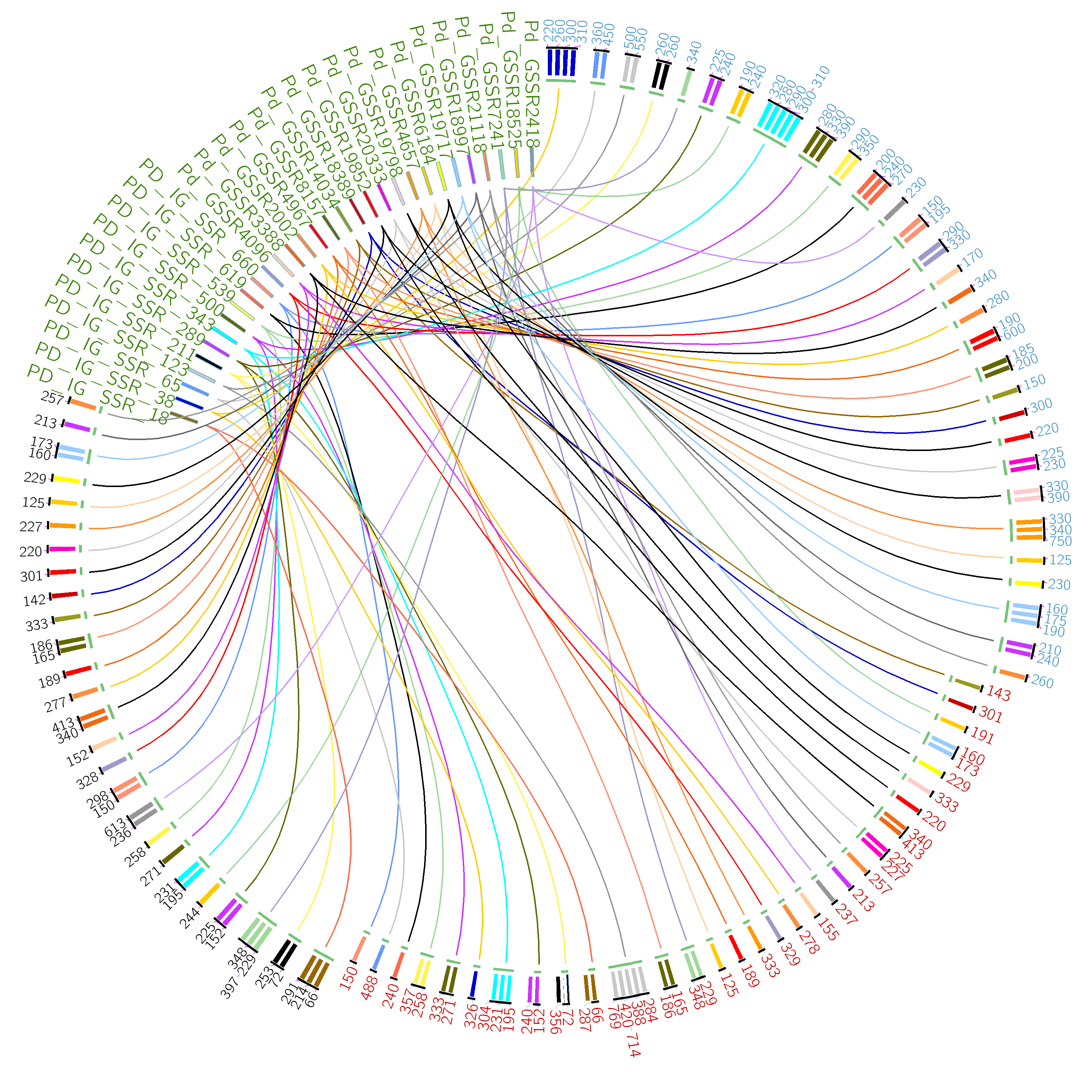
The list of integrated date palm SSRs and Single Nucleotide Polymorphisms (SNPs) were collected from earlier studies (Mokhtar et al., 2016, Al-Dous et al., 2011) and stored in relational database tables of MySQL database. The information of SSR markers was stored in four sub-databases, while, SNP markers was stored in one sub-database. Each SSR or SNP marker was given a unique identifier called Primer-ID. The sub-databases are as follows: